Effects of Temperature Fluctuations on Oscillating Biological Systems

Requires a Wolfram Notebook System
Interact on desktop, mobile and cloud with the free Wolfram Player or other Wolfram Language products.
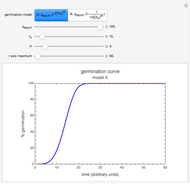
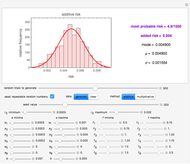
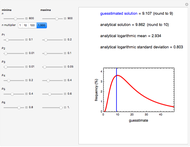
This Demonstration simulates the possible effects of temperature fluctuations on the evolution of oscillating biological populations or systems. It is based on the assumptions that: the amplitude and frequency of the natural oscillation increase with temperature in the pertinent range; the momentary growth rate is the rate at the momentary temperature at the time that corresponds to population size or system state; and the temperature fluctuation pattern can be described by a sum of two sinusoidal terms.
Contributed by: Mark D. Normand and Micha Peleg (December 2011)
Open content licensed under CC BY-NC-SA
Snapshots
Details
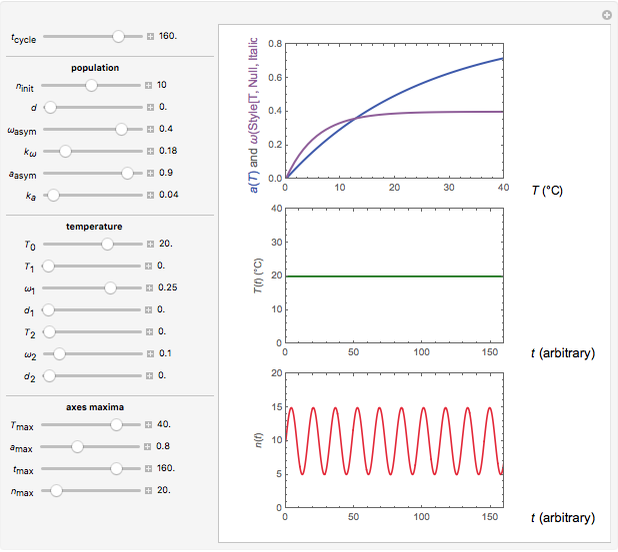
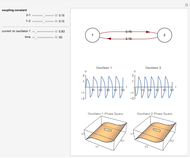
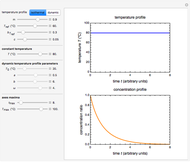
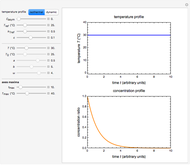
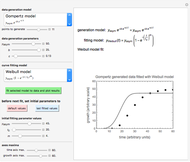
Snapshot 1: oscillation at constant temperature
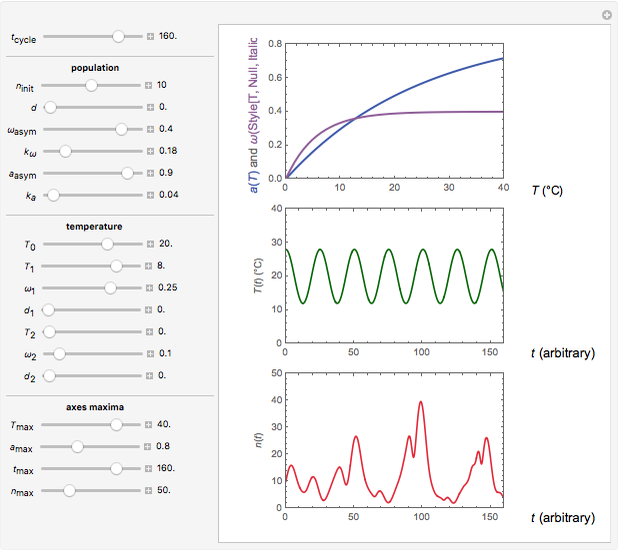
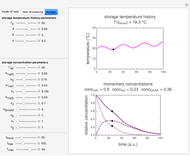
Snapshot 2: oscillation under regular temperature fluctuations
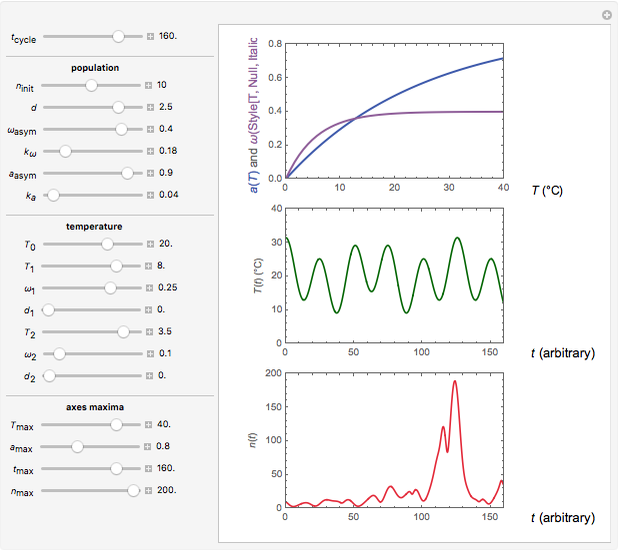
Snapshot 3: oscillation under slightly irregular temperature fluctuations leading to explosion
Snapshot 4: oscillation under slightly irregular temperature fluctuations leading to apparent extinction
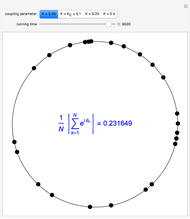
Oscillating biological systems, be they organismic populations or biochemical processes, may be affected by the environment or ambient temperature, which may also fluctuate. This Demonstration illustrates the theoretical possibility that coupling a biological system's oscillations with the temperature's fluctuations can produce apparently chaotic evolution patterns such as "explosions" or a drop to a level that in nature will cause the system's extinction [1]. Such patterns can be generated with simplified models. The one used here is based on the assumption that the biological system's oscillations are single-term sinusoidal and that their amplitude and frequency rise monotonically with temperature in the pertinent temperature range. The environmental or ambient temperature fluctuations are simulated with a two-term sinusoidal model. It is assumed that the momentary rate of change in the biological system's state (e.g., population's size or bioactive compound concentration) under dynamic conditions is the momentary isothermal rate at the momentary temperature, at a time that corresponds to the system's momentary state [2].
The user can choose with sliders the time,  , in arbitrary units; the initial number of organisms or concentration,
, in arbitrary units; the initial number of organisms or concentration, 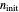 ; the system oscillation's phase,
; the system oscillation's phase,  , in radians; and the parameters
, in radians; and the parameters 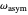 ,
, 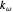 ,
, 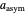 , and
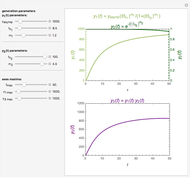
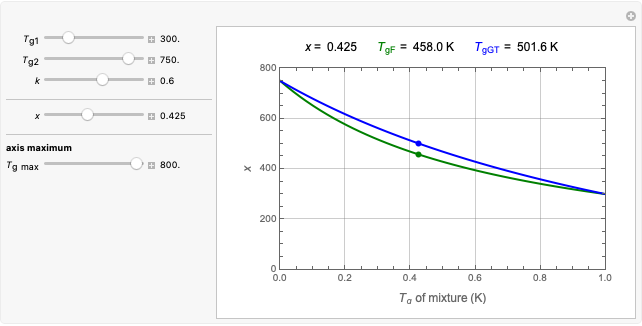
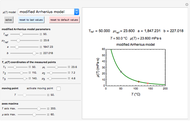
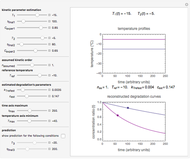
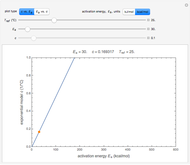
, and 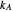 , which determine the oscillation frequency's and amplitude's temperature dependence through the relationships
, which determine the oscillation frequency's and amplitude's temperature dependence through the relationships 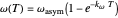 and
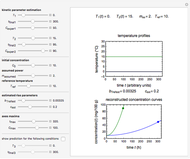
and 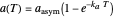 , respectively, which are plotted at the top of the display.
, respectively, which are plotted at the top of the display.
The control parameters of the temperature's fluctuations are 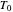 ,
, 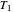 ,
, 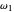 ,
, 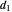 ,
, 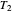 ,
, 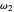 , and
, and 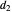 , and they too can be entered with sliders.
, and they too can be entered with sliders. 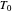 ,
, 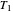 , and
, and 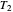 are in degrees C,
are in degrees C, 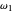 and
and 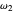 in the arbitrary time unit reciprocal, and
in the arbitrary time unit reciprocal, and 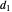 and
and 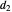 in radians.
in radians.
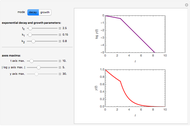
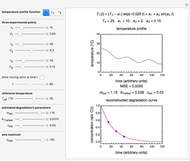
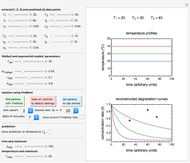
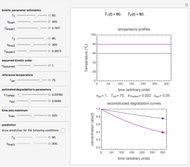
The temperature history, which is plotted in the middle of the graphical display, is generated with the model 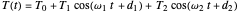 .
.
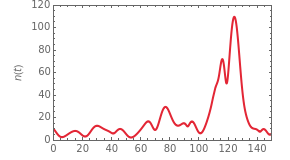
The system's evolution that corresponds to the chosen temperature fluctuation pattern and temperature dependence of the system's own oscillations is plotted at the bottom of the graphical display in the form of the 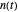 versus
versus  relationship. It is calculated as the numerical solution of the differential equation
relationship. It is calculated as the numerical solution of the differential equation 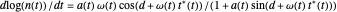 , where
, where 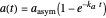 ,
, 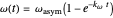 , and
, and 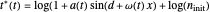 .
.
Except for the time axis range, which is automatically determined by  , the three plots'
, the three plots' 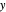 axes maxima can be chosen with sliders.
axes maxima can be chosen with sliders.
Note that setting 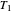 and
and 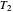 to zero will simulate isothermal conditions at
to zero will simulate isothermal conditions at 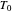 , and that for a larger initial number or a smaller concentration than that allowed by the
, and that for a larger initial number or a smaller concentration than that allowed by the 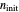 slider, the result can be multiplied or divided by the appropriate factor.
slider, the result can be multiplied or divided by the appropriate factor.
Also note that the purpose of the Demonstration is just to illustrate the theoretical possibility of the emergence of different patterns. Thus, not all the simulated evolution patterns that can be generated with the allowed parameter settings necessarily have a real-life equivalent in the biological world.
References
[1] M. Peleg and M. G. Corradini, "Theoretical Effects of Monotonically Changing and Fluctuating Temperature on Oscillating Biological Systems," Ecological Complexity 7, 2010 pp. 500–505.
[2] M. Peleg, Advanced Quantitative Microbiology for Foods and Biosystems: Models for Predicting Growth and Inactivation, Boca Raton, FL: CRC Press, 2006.
Permanent Citation