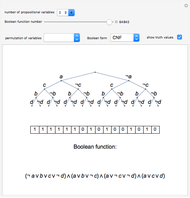
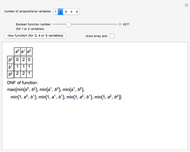
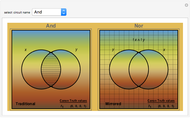
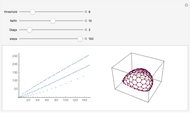
Mutual Information between Boolean Net Regions

Requires a Wolfram Notebook System
Interact on desktop, mobile and cloud with the free Wolfram Player or other Wolfram Language products.
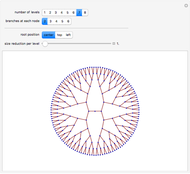
Boolean nets are sequential, dynamical systems that generalize cellular automata. An 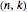 -Boolean net can be represented by a directed graph with
-Boolean net can be represented by a directed graph with  nodes, in which each node is associated with a (different) Boolean function of
nodes, in which each node is associated with a (different) Boolean function of  arguments. All functions are evaluated synchronously, their arguments being identified by the edges incident to each node. This Demonstration supports the creation of Boolean nets with an even number of nodes (
arguments. All functions are evaluated synchronously, their arguments being identified by the edges incident to each node. This Demonstration supports the creation of Boolean nets with an even number of nodes (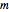 ), partitioned into regions
), partitioned into regions 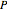 and
and 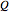 of equal sizes; additionally, you can specify the number of edges connecting
of equal sizes; additionally, you can specify the number of edges connecting 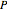 and
and 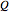 (coupling factor). The main output is an estimate of the asymptotic mutual information between the
(coupling factor). The main output is an estimate of the asymptotic mutual information between the 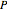 and
and 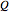 components of the global state as the net evolves.
components of the global state as the net evolves.
Contributed by: Tommaso Bolognesi (May 2018)
Open content licensed under CC BY-NC-SA
Snapshots
Details
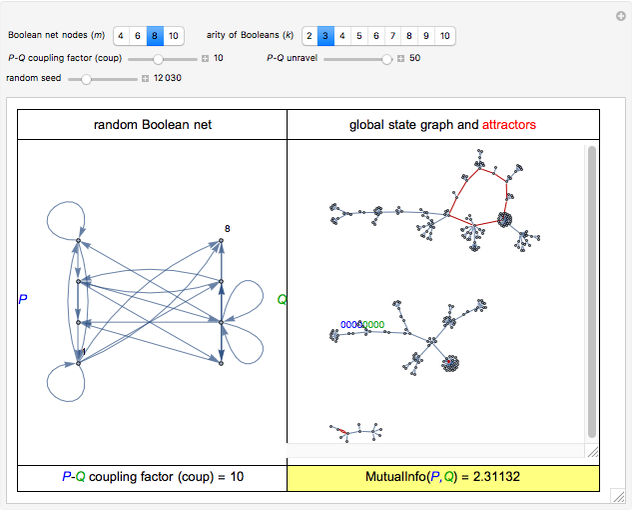
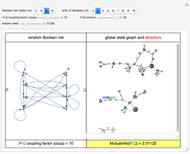
This Demonstration shows the creation of a random Boolean net (RBN) with a user-defined even number of nodes ( ), in which each node is associated with a randomly chosen Boolean function of
), in which each node is associated with a randomly chosen Boolean function of  arguments (
arguments ( ). The node set of the net is split into parts
). The node set of the net is split into parts 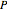 and
and 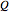 of equal size, and you can assign the
of equal size, and you can assign the 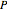 -
-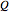 coupling factor "coup", defined as the number of edges that bridge between
coupling factor "coup", defined as the number of edges that bridge between 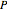 and
and 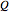 . These edges enable a flow of information between the two net regions. The slider "
. These edges enable a flow of information between the two net regions. The slider "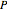 -
-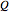 unravel" affects only the layout of the net, providing better visual separation between the nodes in
unravel" affects only the layout of the net, providing better visual separation between the nodes in 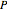 and
and 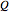 .
.
This Demonstration is concerned with the mutual information 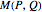 between the states of the
between the states of the 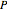 and
and 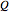 net components as the net evolves; these states are
net components as the net evolves; these states are 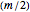 -tuples of bits and are seen as random variables, also denoted by
-tuples of bits and are seen as random variables, also denoted by 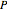 and
and 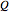 . The mutual information
. The mutual information  between two random variables measures the amount of information that one variable provides about the other and is defined in terms of their joint and marginal distributions. When two variables are independent, their mutual information is 0.
between two random variables measures the amount of information that one variable provides about the other and is defined in terms of their joint and marginal distributions. When two variables are independent, their mutual information is 0.
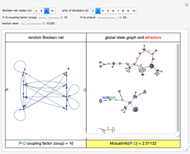
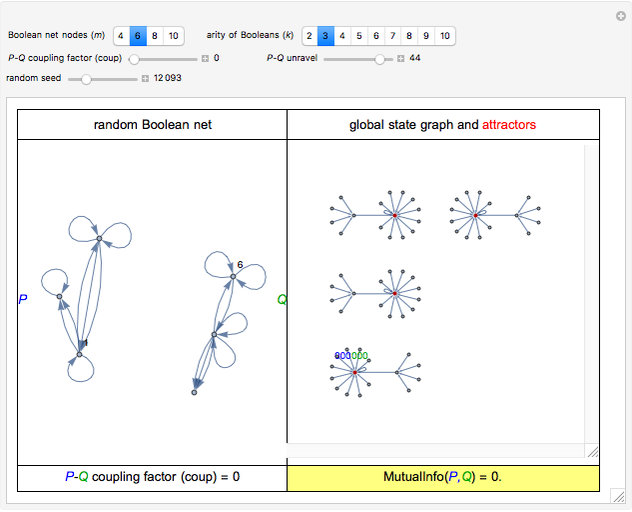
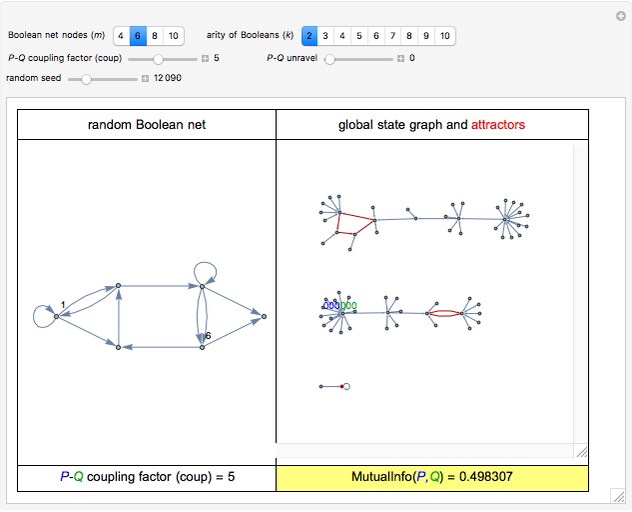
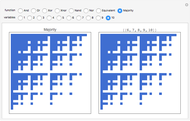
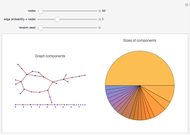
For computational efficiency, this Demonstration adopts an algorithm that provides an approximate but sufficiently accurate estimate of the asymptotic value of 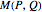 or of its average in the case of periodic fluctuation. The approximation is obtained by analyzing the global state graph of the net and its attractors, depicted in the right subpanel. Each global state is an
or of its average in the case of periodic fluctuation. The approximation is obtained by analyzing the global state graph of the net and its attractors, depicted in the right subpanel. Each global state is an 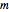 -tuple of bits (only tuple
-tuple of bits (only tuple  is actually shown in the graph) that we conceive as split into two tuples of equal length, colored in blue and green and corresponding to random variables
is actually shown in the graph) that we conceive as split into two tuples of equal length, colored in blue and green and corresponding to random variables 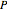 and
and 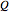 . We assume an initial uniform distribution of all global states. Under this distribution, variables
. We assume an initial uniform distribution of all global states. Under this distribution, variables 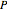 and
and 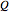 are initially independent: any value of
are initially independent: any value of 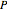 is equally likely to be paired with any value of
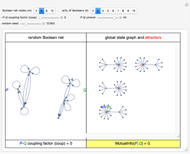
is equally likely to be paired with any value of 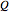 . If coup
. If coup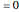 ,
, 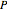 and
and 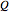 remain independent as the net evolves, the theoretical value of the corresponding asymptotic mutual information is
remain independent as the net evolves, the theoretical value of the corresponding asymptotic mutual information is 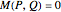 . Interestingly, the two variables remain independent also when coup assumes its maximum possible value
. Interestingly, the two variables remain independent also when coup assumes its maximum possible value  , yielding again
, yielding again 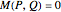 . (Occasionally, for these two extreme coup values the
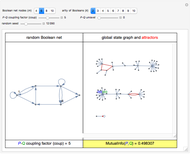
. (Occasionally, for these two extreme coup values the 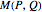 shown in the lower-right slot of the panel may slightly depart from 0, due to the approximation procedure.) Conversely, nonzero values of coup typically yield nonzero asymptotic values of
shown in the lower-right slot of the panel may slightly depart from 0, due to the approximation procedure.) Conversely, nonzero values of coup typically yield nonzero asymptotic values of 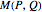 . On average,
. On average,  would describe a reversed parabola as coup moves from 0 to
would describe a reversed parabola as coup moves from 0 to  .
.
References
[1] S. Kauffman, "Homeostasis and Differentiation in Random Genetic Control Networks," Nature, 224, 1969 pp. 177–178. doi:10.1038/224177a0.
[2] S. Kauffman, At Home in the Universe: The Search for Laws of Self-Organization and Complexity, New York: Oxford University Press, 1995.
[3] D. Balduzzi and G. Tononi, "Integrated Information in Discrete Dynamical Systems: Motivation and Theoretical Framework," PLOS Computational Biology, 4(6), 2008 e1000091. doi:10.1371/journal.pcbi.1000091.
Permanent Citation