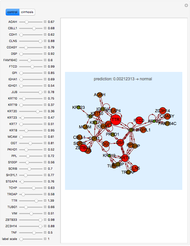
Ramachandran Analysis of Protein Backbone Dihedral Angles

Requires a Wolfram Notebook System
Interact on desktop, mobile and cloud with the free Wolfram Player or other Wolfram Language products.
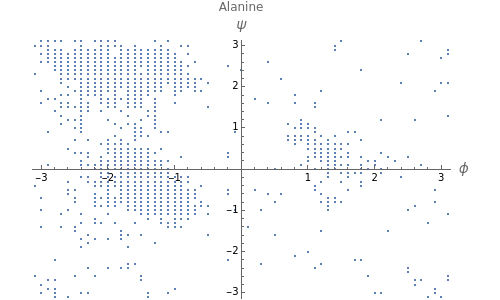
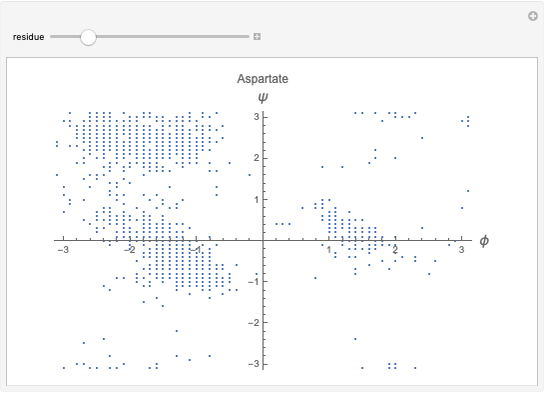
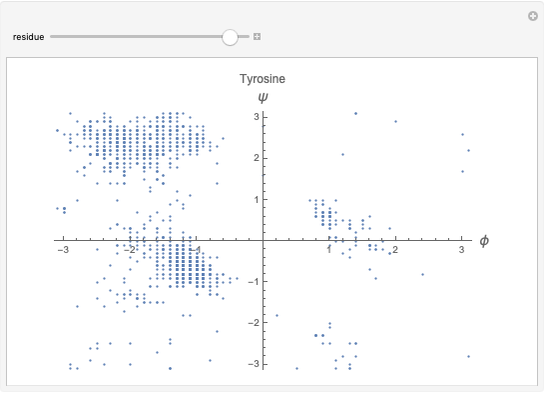
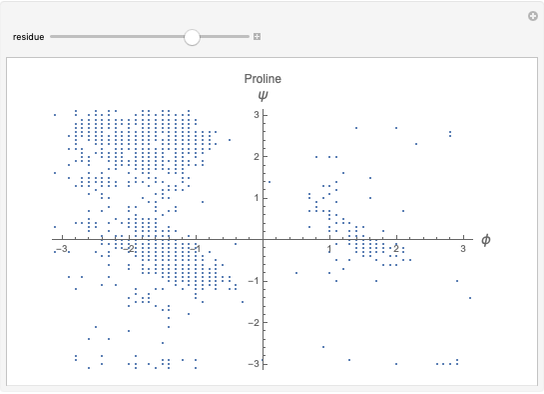
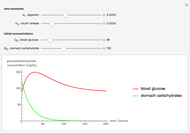
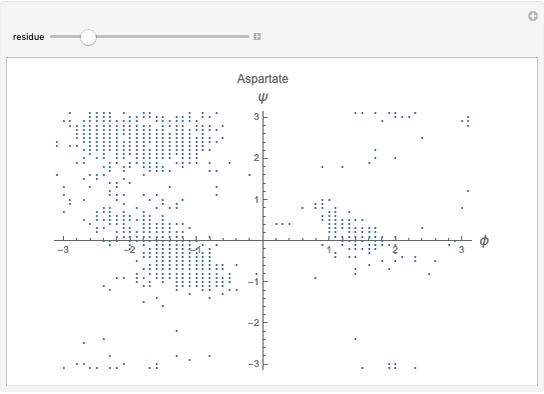
This Demonstration shows the distribution of backbone dihedral angles for each amino acid in a representative set of human proteins tabulated in Mathematica via the ProteinData command. Available conformations are restricted due to steric clashes of the side chains; Ramachandran analysis reveals that certain conformations are preferentially accessible to amino acids.
Contributed by: Daniel Barr (January 2013)
Open content licensed under CC BY-NC-SA
Snapshots
Details
Ramachandran analysis demonstrates that only a subset of the possible values for the backbone dihedral angle in a protein are actually observed in protein structure due to steric repulsions between amino acid side chains. The  and
and 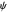 angles measure rotations around the N-C
angles measure rotations around the N-C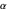 and C
and C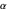 -C bonds, respectively, and are by convention plotted over a range from
-C bonds, respectively, and are by convention plotted over a range from 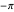 to
to 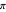 (corresponding to
(corresponding to 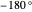 to
to 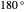 ).
).
Most amino acids fall into well-defined regions of the Ramachandran plot (see, e.g. "Ramachandran Plot"). Due to their unique side chains, glycine and proline show significant population of conformations in the traditionally forbidden regions of the Ramachandran plot.
Permanent Citation