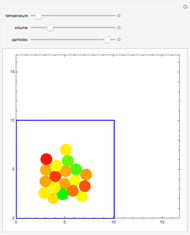
Molecular Dynamics of Lennard-Jones Particles Using the Velocity Verlet Algorithm

Requires a Wolfram Notebook System
Interact on desktop, mobile and cloud with the free Wolfram Player or other Wolfram Language products.
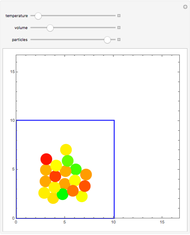
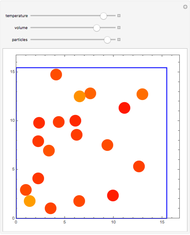
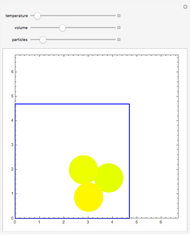
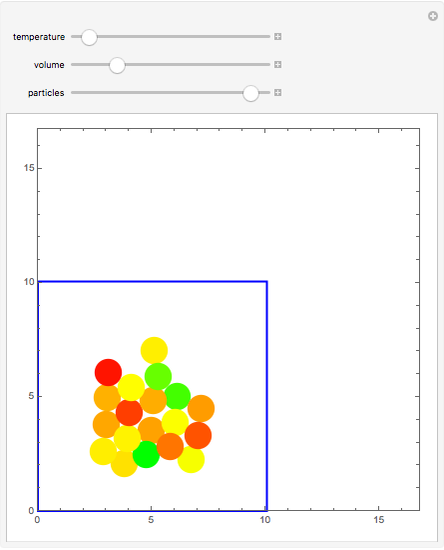
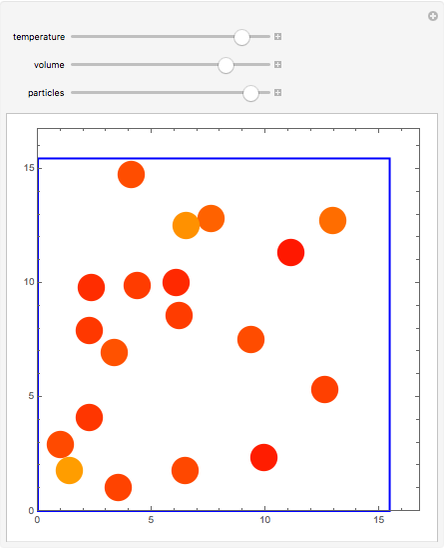
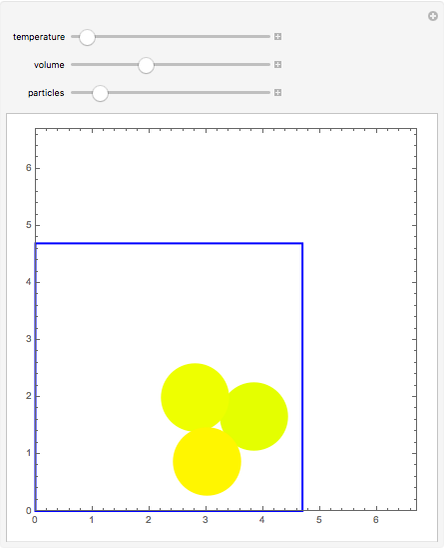
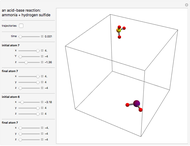
Molecular dynamics is a computer simulation technique that follows the time evolution of a set of interacting atoms or molecules by integrating their equations of motion. This Demonstration uses molecular dynamics and the velocity Verlet algorithm to simulate the motion of particles interacting under the Lennard–Jones 6-12 potential. You can vary the temperature, volume, and number of particles to change from the formation of clusters to liquid-like behavior. The color of each particle depends on its speed.
Contributed by: José Luis Gómez-Muñoz (March 2011)
Open content licensed under CC BY-NC-SA