Prediction of Isothermal Degradation by the Endpoints Method

Requires a Wolfram Notebook System
Interact on desktop, mobile and cloud with the free Wolfram Player or other Wolfram Language products.
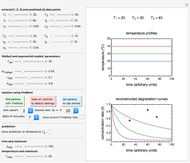
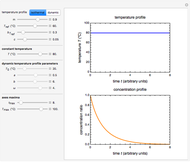
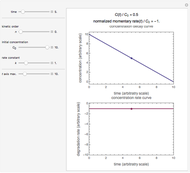
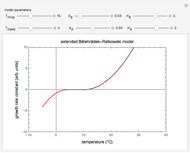
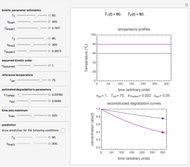
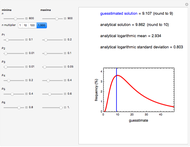
Theoretically, when a degradation reaction is known to follow known fixed-order kinetics, its parameters can be determined from two final concentration ratios at two constant temperatures by numerically solving two simultaneous rate equations. Such a solution, however, may require fairly accurate initial guesses that are difficult to assign intuitively. Nevertheless, they can be estimated by manually matching the two experimental points (shown in red and blue) with their corresponding degradation curves using Mathematica’s Manipulate function. Once found, they can be used either as initial guesses or as parameters themselves. This is shown with user-entered endpoints assuming first-order or approximately first-order kinetics and showing how the retrieved parameters can be used to predict the concentration ratio at different temperatures after different times.
Contributed by: Mark D. Normandand Micha Peleg (April 2015)
Open content licensed under CC BY-NC-SA
Snapshots
Details
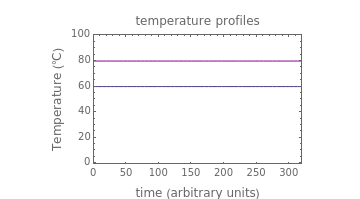
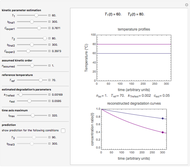
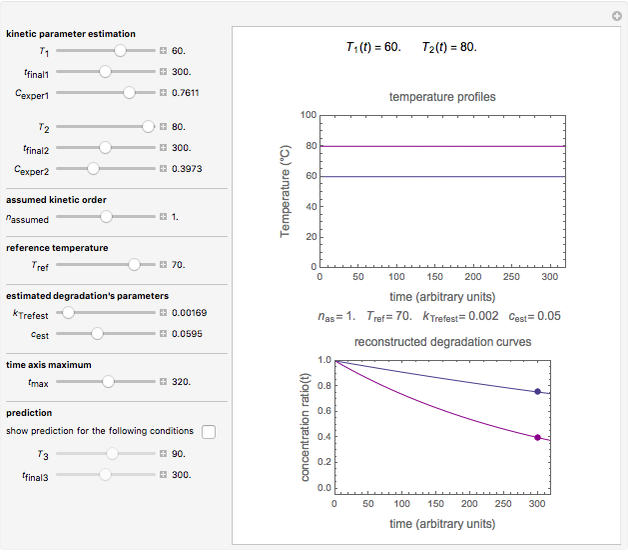
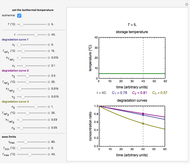
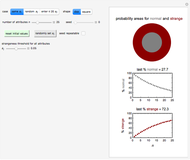
Snapshot 1: the two concentration ratios shown in the thumbnail matched manually with the corresponding degradation curves, assuming first-order kinetics ( )
)
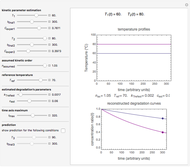
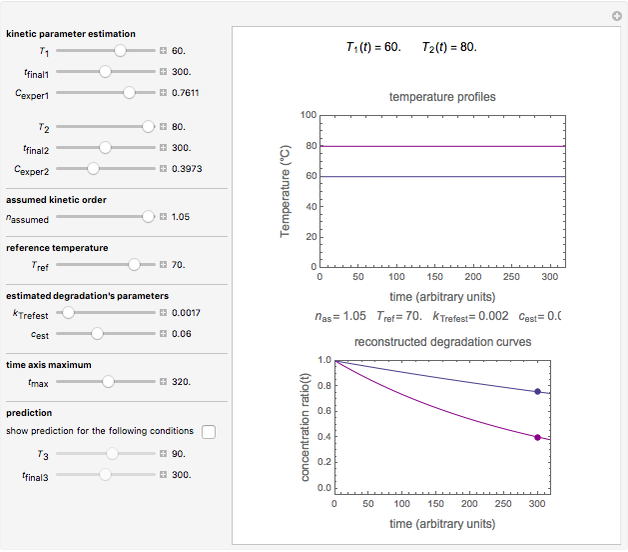
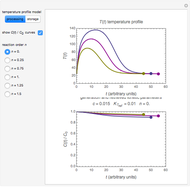
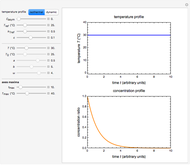
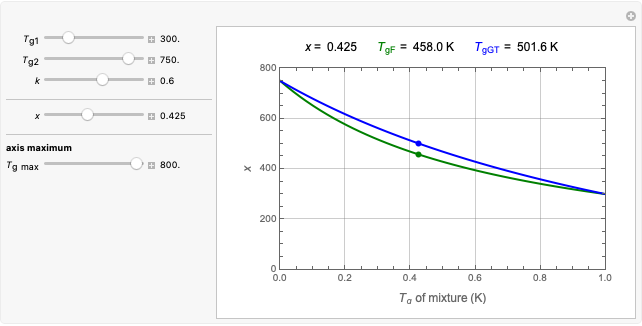
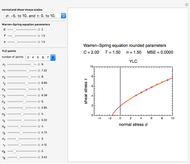
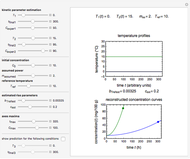
Snapshot 2: two concentration ratios at different times matched manually with the corresponding degradation curves, assuming approximate first-order kinetics with 
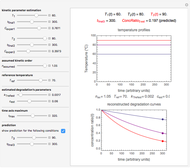
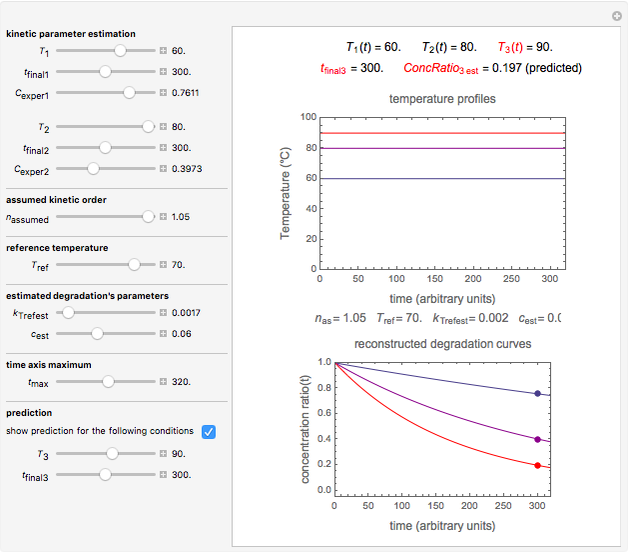
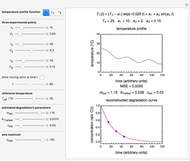
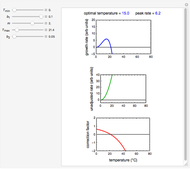
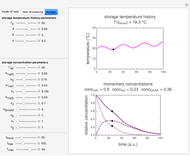
Snapshot 3: predicting isothermal degradation (red curve) at time  and temperature
and temperature  (red point) from the matched points in Snapshot 2
(red point) from the matched points in Snapshot 2
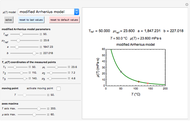
Consider a degradation reaction of a heat-labile biological compound, for example, a vitamin or pigment, following assumed fixed-order kinetics ( ), where the temperature-dependence of the rate constant follows the exponential form
), where the temperature-dependence of the rate constant follows the exponential form  , where
, where  is the rate constant at temperature
is the rate constant at temperature  . As has already been shown, this simpler model can replace the traditional Arrhenius equation without sacrificing the fit [1].
. As has already been shown, this simpler model can replace the traditional Arrhenius equation without sacrificing the fit [1].
When the compound is exposed to two constant temperatures  and
and  for sufficiently long times so that the come-up and cooling times' effects are negligible, the two corresponding degradation curves can be modelled by
for sufficiently long times so that the come-up and cooling times' effects are negligible, the two corresponding degradation curves can be modelled by  and
and  . The final concentrations
. The final concentrations  and
and  are the solutions of these two rate equations for times
are the solutions of these two rate equations for times  and
and  , respectively. This gives two equations
, respectively. This gives two equations  and
and  with two unknowns,
with two unknowns,  and
and  .
.
If the final concentration ratios have actually been measured, then one can solve these two equations simultaneously using FindRoot and extract the values of  and
and  [2]. Once calculated, these parameters can be used to reconstruct the entire degradation curves of the two temperatures and predict other temperature histories, isothermal or nonisothermal.
[2]. Once calculated, these parameters can be used to reconstruct the entire degradation curves of the two temperatures and predict other temperature histories, isothermal or nonisothermal.
In this Demonstration, enter the two temperatures  and
and  , corresponding concentration ratios
, corresponding concentration ratios  and
and  at chosen times
at chosen times  and
and  , the assumed reaction order
, the assumed reaction order  (
( ), and chosen reference temperature
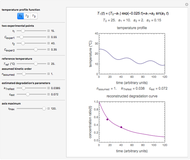
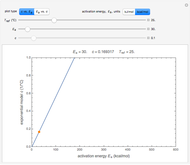
), and chosen reference temperature  between or equal to one of the two chosen temperatures. Once these are set and displayed, one tries to match the two shown degradation curves with the two chosen points by moving the
between or equal to one of the two chosen temperatures. Once these are set and displayed, one tries to match the two shown degradation curves with the two chosen points by moving the  and
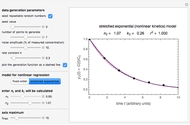
and  sliders. The match can be improved by adjusting the numerical values of these two parameters displayed to the right of the sliders. When a satisfactory match is obtained, the corresponding parameters' values, which are displayed on top of the bottom plot, can be used as the initial guesses for solving the equations or considered as estimates of the parameters' values themselves. In that case, by checking the "show prediction for the following conditions" box, you can estimate and display the concentration ratio at any temperature and time entered by the
sliders. The match can be improved by adjusting the numerical values of these two parameters displayed to the right of the sliders. When a satisfactory match is obtained, the corresponding parameters' values, which are displayed on top of the bottom plot, can be used as the initial guesses for solving the equations or considered as estimates of the parameters' values themselves. In that case, by checking the "show prediction for the following conditions" box, you can estimate and display the concentration ratio at any temperature and time entered by the  and
and  sliders.
sliders.
Not all time-temperature-concentration ratio entries allowed by the program must have two matching degradation curves. Some entries may reflect experimental errors, a reaction kinetic order outside the permitted range, or nonlinear kinetics.
References
[1] M. Peleg, M. D. Normand, and M. G. Corradini, "The Arrhenius Equation Revisited," Critical Reviews in Foods Science and Nutrition, 52(9), 2012 pp. 830–851. doi:10.1080/10408398.2012.667460.
[2] M. Peleg, M. D. Normand, and A. D. Kim, "Estimating Thermal Degradation Kinetics Parameters from the Endpoints of Non-isothermal Heat Processes or Storage," Food Research International, 66, 2014 pp. 313–324. doi:10.1016/j.foodres.2014.10.003.
Permanent Citation