Estimation of Time to Excessive Microbial Count

Requires a Wolfram Notebook System
Interact on desktop, mobile and cloud with the free Wolfram Player or other Wolfram Language products.
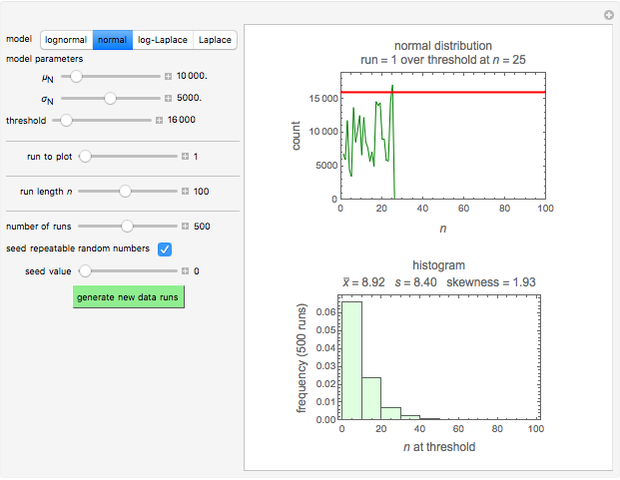
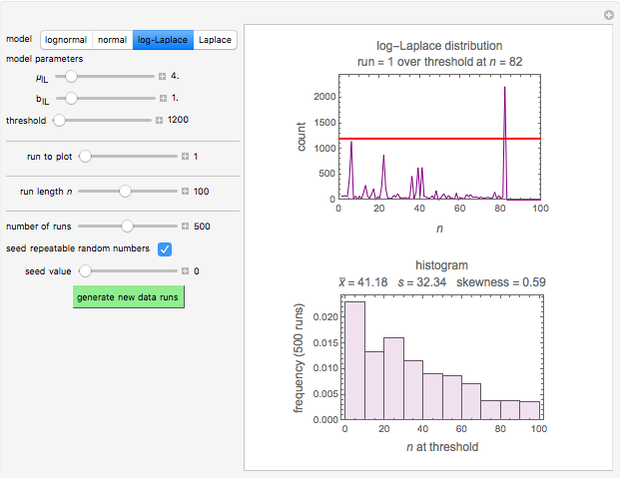
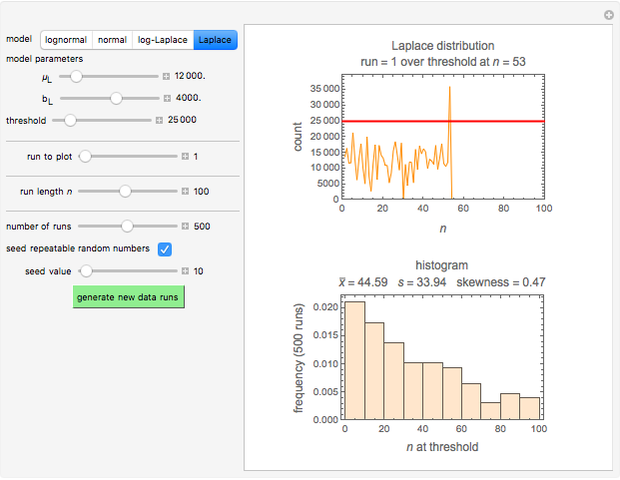
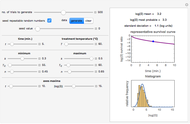
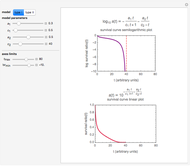
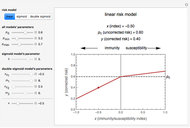
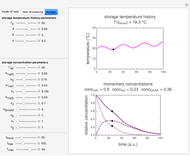
Many microbial count records in foods and water resemble a random time series. The distribution of the entries allows you to estimate the frequency of future counts exceeding a level deemed undesirable or dangerous. This Demonstration lets you generate Monte Carlo simulations of such counts using the lognormal, normal, log-Laplace, or Laplace distribution function as a model, with hypothetical or experimentally determined parameters, and record the times at which a specified threshold has been exceeded. You can use the distribution and histogram of these times to assess how soon an excessive count might be encountered if conditions remain unchanged.
Contributed by: Mark D. Normand and Micha Peleg (November 2012)
Open content licensed under CC BY-NC-SA
Snapshots
Details
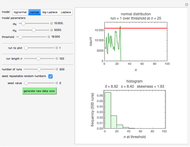
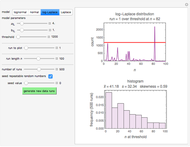
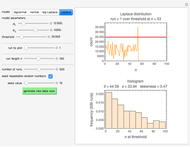
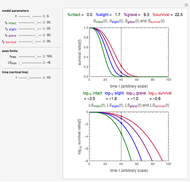
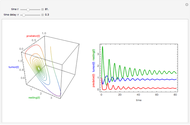
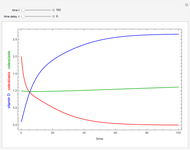
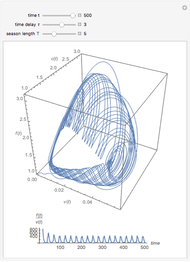
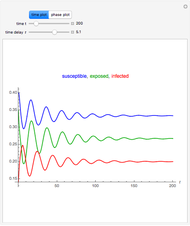
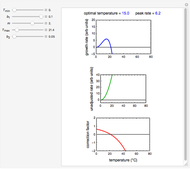
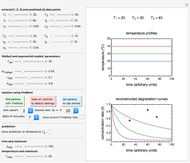
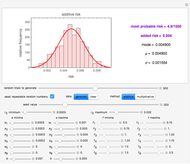
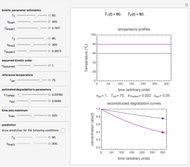
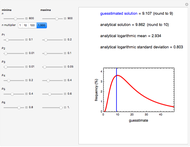
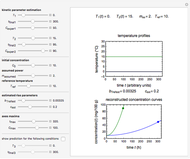
Snapshots 1, 2, 3: simulated fluctuating microbial counts having a Laplace, normal, or log-Laplace distribution, with the chosen parameters and a histogram of the times to exceed the specified upper threshold
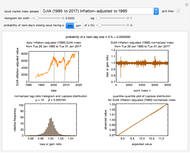
Microbial count records encountered in foods and water frequently resemble a randomly fluctuating time series with the entries having a characteristic underlying distribution. Provided that the entries are independent and have no trend or periodicity, their distribution allows you to estimate the probability, or future frequency, of encountering a count exceeding an undesirable threshold [1–3].
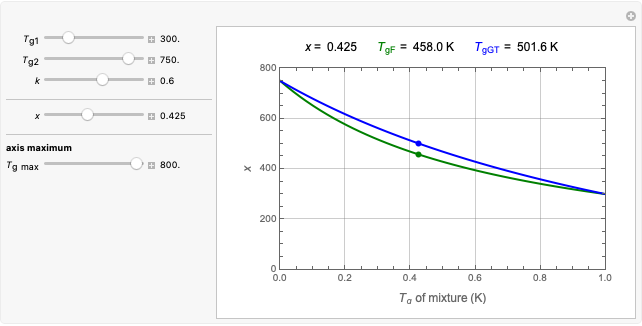
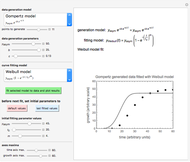
When the counts are taken at fixed intervals, their distribution also allows you to estimate the most probable first time of such an occurrence. This Demonstration generates Monte Carlo simulations of random microbial count series using the lognormal, normal, log-Laplace, or Laplace distributions as the fluctuation model [2, 3]. If the threshold is exceeded, the run terminates. The program keeps every run, which you can view individually with the chosen threshold superimposed. The program also creates and displays a histogram of the times at which the chosen threshold has been exceeded and shows their calculated mean, standard deviation, and skewness.
The counts' distribution function is selected with a setter bar and you can vary its parameters, whether they are hypothetical or obtained from experimental data. You can also change the threshold level, the run to plot, the runs' length, the number of runs to be generated, and the seed, in case you want to generate reproducible count records.
If at a particular setting all the runs start with a count already exceeding the threshold, the plots are replaced by a message in red alerting you to this. A message is displayed in blue when none of the generated counts exceeds the threshold in any of the runs.
References
[1] M. Peleg and J. Horowitz, "On Estimating the Probability of Aperiodic Outbursts of Microbial Populations from Their Fluctuating Counts," Bulletin of Mathematical Biology, 62(1), 2000 pp. 17–35. link.springer.com/content/pdf/10.1006%2 Fbulm .1999.0112.
[2] M. G. Corradini, M. D. Normand, A. Nussinovitch, J. Horowitz, and M. Peleg, "Estimating the Frequency of High Microbial Counts in Commercial Food Products Using Various Distribution Functions," Journal of Food Protection, 64(5), 2001 pp. 674–681. www.ncbi.nlm.nih.gov/pubmed/11347999.
[3] M. Peleg, Advanced Quantitative Microbiology for Food and Biosystems: Models for Predicting Growth and Inactivation, Boca Raton, FL: CRC Press, 2006.
Permanent Citation
"Estimation of Time to Excessive Microbial Count"
http://demonstrations.wolfram.com/EstimationOfTimeToExcessiveMicrobialCount/
Wolfram Demonstrations Project
Published: November 29 2012